Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
Por um escritor misterioso
Descrição
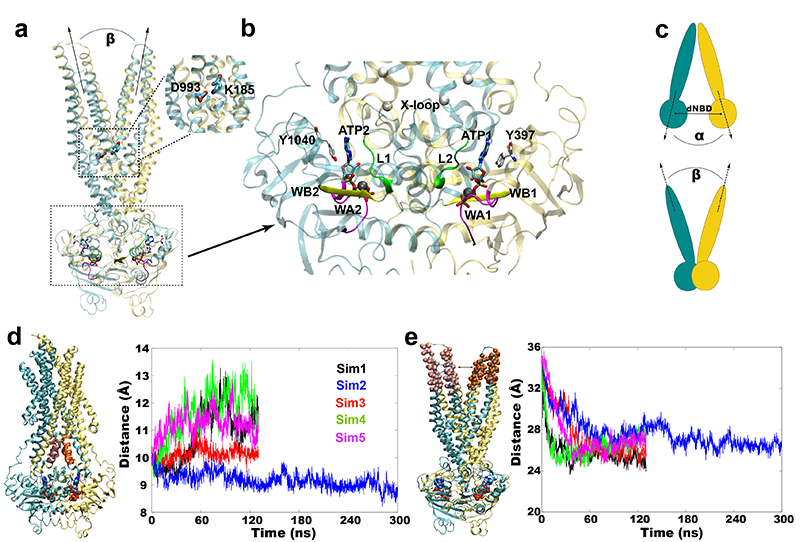
Novel Energy Transduction in P-glycoprotein
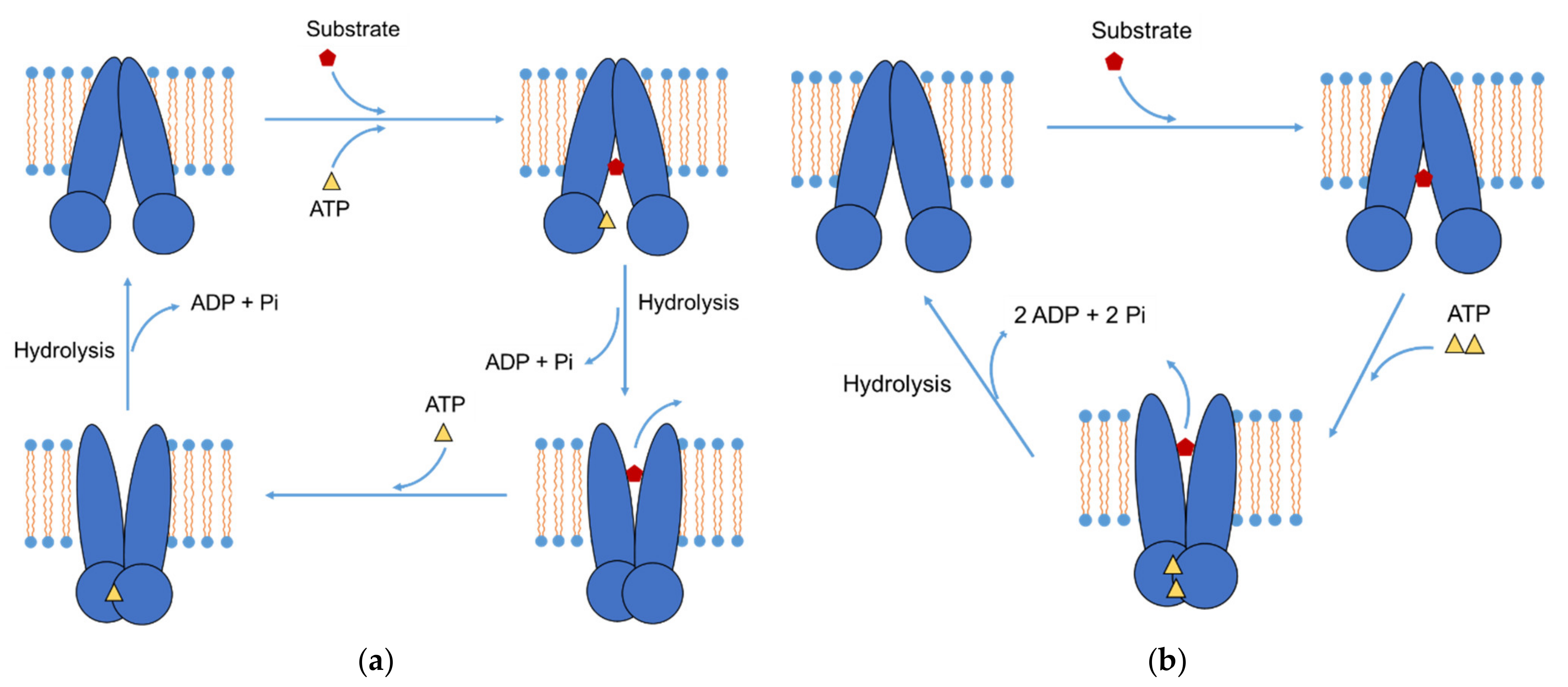
Pharmaceutics, Free Full-Text
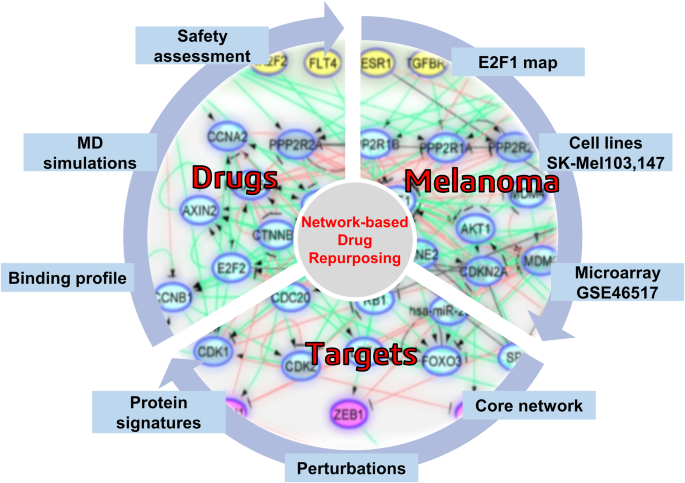
Logic-based modeling and drug repurposing for the prediction of novel therapeutic targets and combination regimens against E2F1-driven melanoma progression, BMC Chemistry

Multisite model for P-glycoprotein drug binding. MOLCAD representation
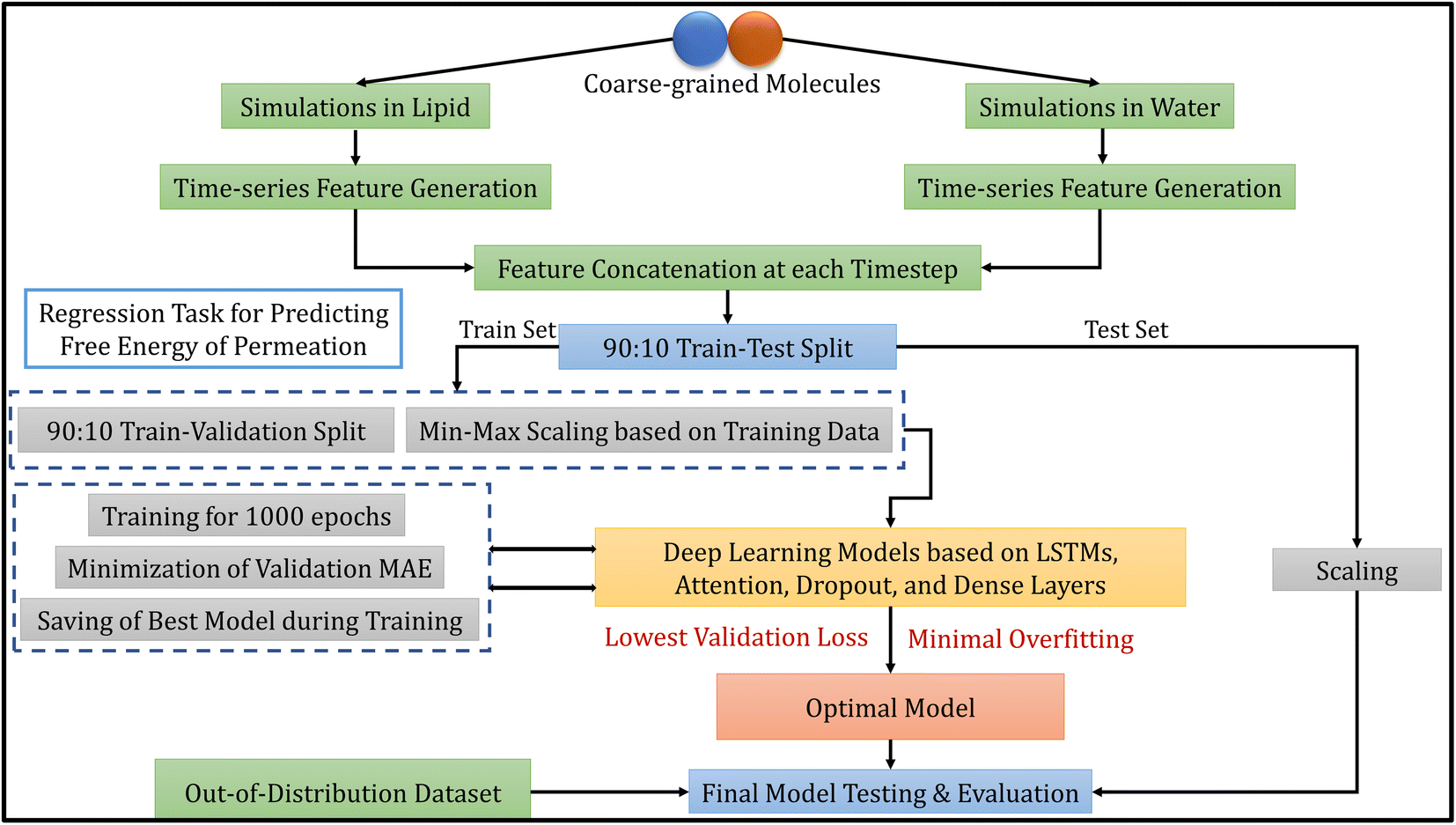
Deep learning models for the estimation of free energy of permeation of small molecules across lipid membranes - Digital Discovery (RSC Publishing) DOI:10.1039/D2DD00119E

Combining machine‐learning and molecular‐modeling methods for drug‐target affinity predictions - Perez‐Lopez - 2023 - WIREs Computational Molecular Science - Wiley Online Library

Machine learning approaches and their applications in drug discovery and design - Priya - 2022 - Chemical Biology & Drug Design - Wiley Online Library

Computational and artificial intelligence-based approaches for drug metabolism and transport prediction: Trends in Pharmacological Sciences

Development of an In Silico Prediction Model for P-glycoprotein Efflux Potential in Brain Capillary Endothelial Cells toward the Prediction of Brain Penetration

Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
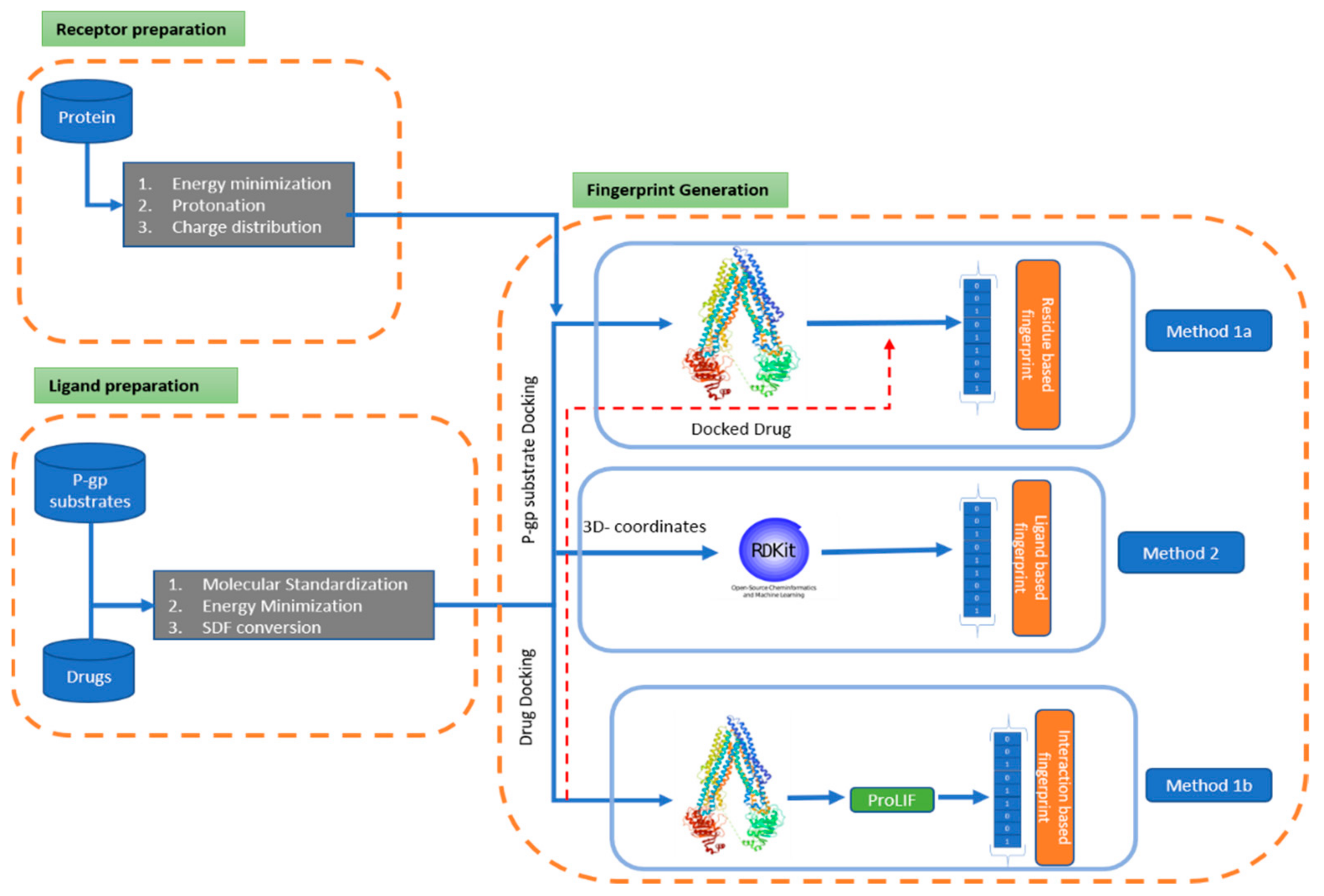
IJERPH, Free Full-Text
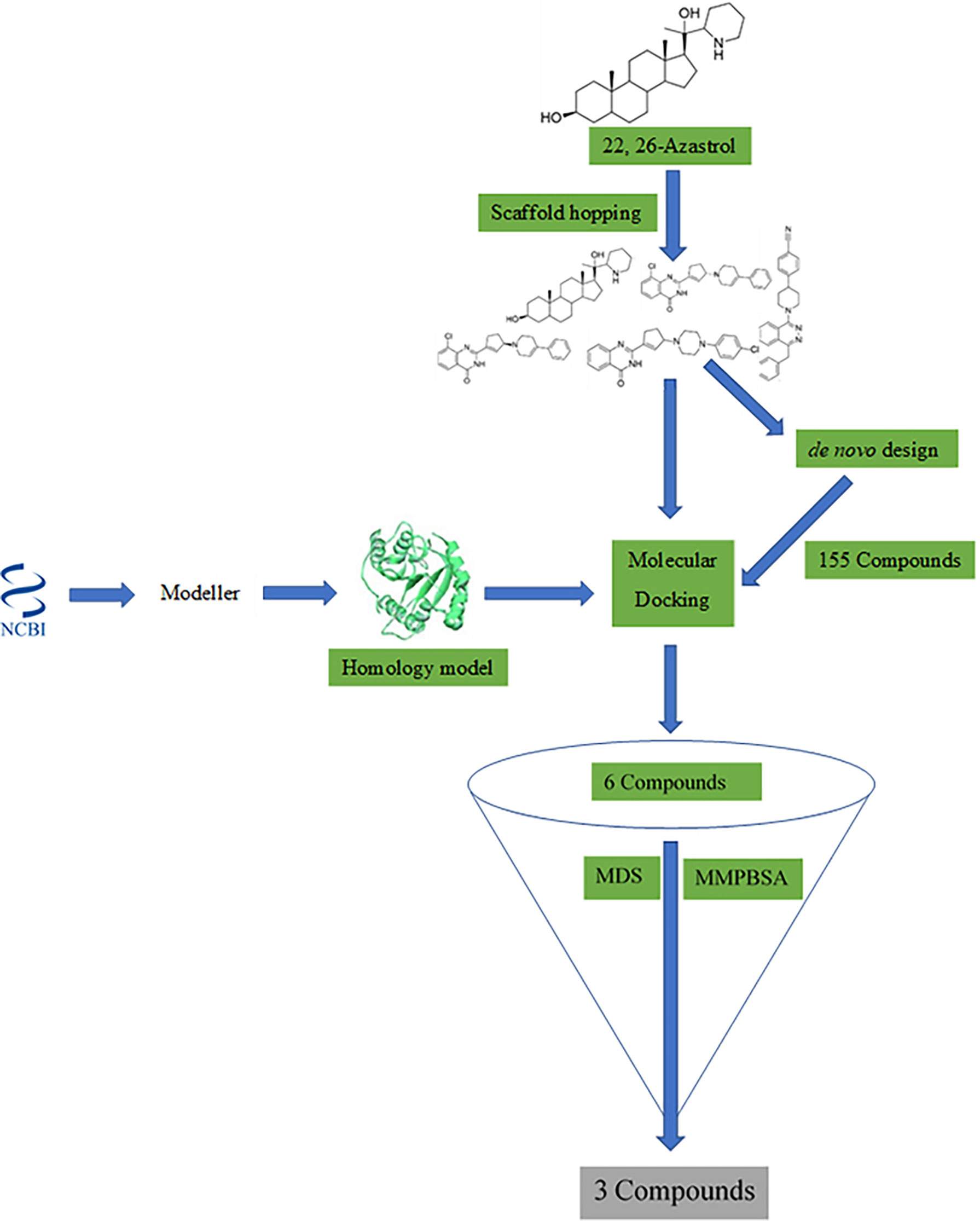
Frontiers Homology Modeling, de Novo Design of Ligands, and Molecular Docking Identify Potential Inhibitors of Leishmania donovani 24-Sterol Methyltransferase

Predicting drug metabolism and pharmacokinetics features of in-house compounds by a hybrid machine-learning model - ScienceDirect

P-glycoprotein Substrate Models Using Support Vector Machines Based on a Comprehensive Data set

P-gp substrate probes (A), known P-gp substrates and an MRP2 inhibitor
de
por adulto (o preço varia de acordo com o tamanho do grupo)